|
Ontozoo is a web page that describes all the ontology software programs primarily developed in the He Group.
Onto-animal tools are a package of web-based ontology tools developed to support efficient and integrated ontology development and application. This package of tools includes OntoFox, Ontodog, Ontorat, Ontobee, Ontobeep, Ontobat, and GOfox. Each tool has specific functions; together, these tools support the extraction of a single or subset of terms and community views from existing ontologies, generation and editing of ontology terms, query and visualization of ontology terms, comparison among ontologies, and instance-level data representation and analysis. Based on the Web Ontology Language (OWL) and Semantics Web technologies, these tools have widely been used by thousands of ontology developers in over 20 communities.
Table of contents:
- Introduction of Onto-animal tools
- OntoFox
- Ontobee
- Ontobeep
- Ontodog
- Ontorat
- Ontobat
- GOfox
- BFOConvert
- Hegroup RDF triple store
- Acknowledgements
A. Introduction of Onto-animal tools:
Biological/biomedical ontologies are sets of computer- and human-interpretable terms and relations that represent entities and their relations in the biological/biomedical world. Biomedical ontologies have emerged as a major tool for the integration and analysis of the large amounts of heterogeneous biological data available in the post-genomics era.
To support ontology development and applications, we have developed a collection of “Onto-animal” tools, including OntoFox, Ontodog, Ontorat, Ontobee, Ontobeep, and Ontobat. The back-end of these Onto-animal tools is the He group’s RDF triple store, which has become the default ontology RDF triple store for the Open Biological and Biomedical Ontologies (OBO) Foundry ontologies.
The Onto-animal tools are summarized in the following figure and described in the following sections:
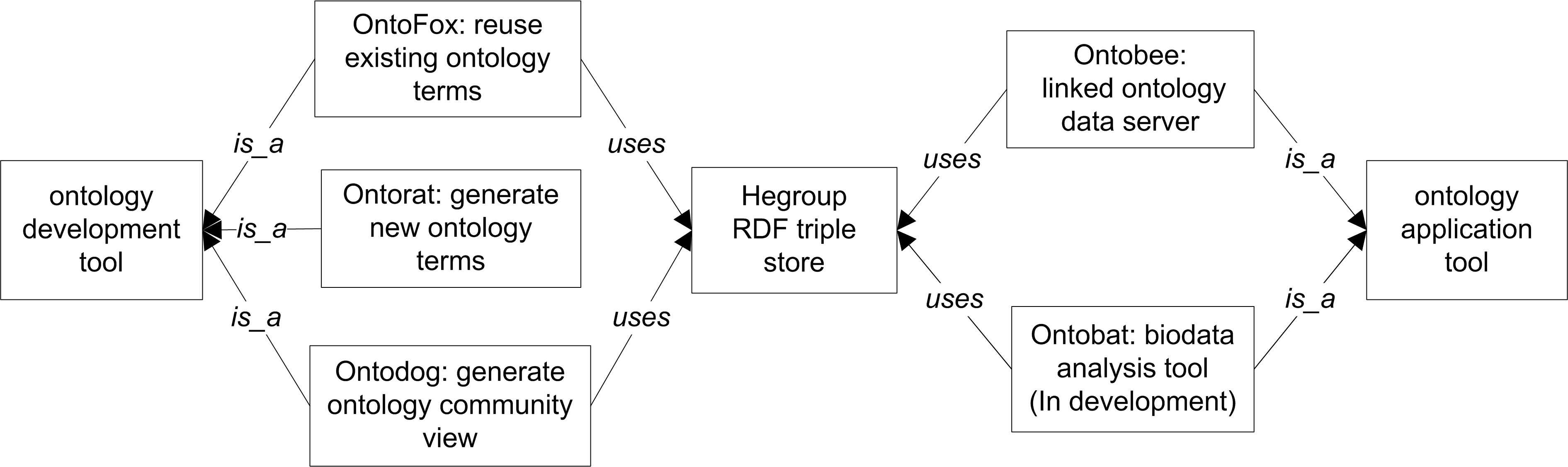
Reference: He Y, Zheng J, Lin Y. Onto-animal tools for reusing ontologies, generating and editing ontology terms, and dereferencing ontology terms.International Conference on Biomedical Ontologies (ICBO-2015). Lisbon, Portugal, July 27-30, 2015. 2-page short paper for poster presentation and flask talk. [Flash Talk] [Poster] [Short paper].
B. OntoFox: Extract ontology terms and axioms (http://ontofox.hegroup.org/)
OntoFox: An Ontology tool that Fetches ontology terms and axioms. OntoFox is a web-based system to support ontology reuse.
Reusing a portion of an existing ontology is often required in the ontology development process. After a user provides a single or a set of terms of interest, OntoFox is able to fetch selected classes, properties, annotations, and their related terms from source ontologies and save the results in the OWL format (Xiang et al., 2010). OntoFox implements the Minimum Information to Reference an External Ontology Term (MIREOT) strategy by extracting minimum information of requested terms. In addition, by providing different options, OntoFox can extract different levels of intermediate terms between the required terms and a chosen higher level or top term. Inspired by existing ontology modularization techniques. OntoFox also implements a new SPARQL-based ontology term extraction algorithm that extracts all terms and axioms related to a given set of user-provided terms.
Reference: Xiang Z, Courtot M, Brinkman RR, Ruttenberg A, He Y. OntoFox: web-based support for ontology reuse. BMC Research Notes. 2010, 3:175. [PMID: 20569493]
C. Ontobee: Linked data server for web displaying and dereferencing ontology terms (http://www.ontobee.org)
Ontobee is an ontology browser and a linked ontology data server for dereferencing ontology terms. To date, there are >100 ontologies listed on Ontobee. Ontobee loads individual page for each term in an ontology. All related information of a single term, such as label, definition, synonyms, superclass hierarchy, logical axioms, and term usage by other ontologies is displayed. In addition, for each ontology, Ontobee generates statistics with counts of classes, object properties, annotation properties, and datatype properties based on term ontology prefixes. Furthermore, Ontobee automatically provides an Excel document listing all terms in an ontology. Ontobee provides ontology term search and SPARQL query service supported by the He group triple store. Ontobee is a de facto search engine for OBO Foundry ontologies.
Reference: Xiang Z, Mungall C, Ruttenberg A, He Y. Ontobee: A Linked Data Server and Browser for Ontology Terms. Proceedings of the 2nd International Conference on Biomedical Ontologies (ICBO), July 28-30, 2011, Buffalo, NY, USA. Pages 279-281. URL: http://ceur-ws.org/Vol-833/paper48.pdf.
D. Ontobeep: Ontology comparison (http://www.ontobee.org/ontobeep/)
Ontobeep is an ontology comparison program. Ontobeep can be used to compare different ontologies by aligning them from the roots of these ontologies. The alignment identifies common terms existing in two or three ontologies. Ontobeep also provides a statistic report of the alignment analysis. Ontobeep may be utilized to detect inconsistency and term duplication in one or more ontologies.
Reference:
E. Ontodog: Generate ontology community view (http://ontodog.hegroup.org/)
Similar to OntoFox, the web-based Ontodog program is able to extract a subset of ontology terms and axioms. Unlike OntoFox, Ontodog includes two unique features. First, Ontodog allows the generation of an ontology community view, which we have defined as “the whole or a subset of the source ontology with user-specified annotations including user preferred labels”. Second, Ontodog uses Excel input files to identify which terms to retrieve and to add user-specified annotations. Excel templates are also provided for easy implementation.
Reference: Zheng J, Xiang Z, Stoeckert Jr. CJ, He Y. Ontodog: a web-based ontology community view generation tool. Bioinformatics. 2014; doi: 10.1093/bioinformatics/btu008. [PDF] PMID: 24413522.
F. Ontorat: Adding new terms and new axioms to an ontology based on design pattern (http://ontorat.hegroup.org)
Ontorat: An ontology tool that supports Ontology representation of axioms using templates and design patterns.
The Ontorat program automatically generates and edits ontology terms and axioms and provides term annotations. Ontorat uses reusable ontology design patterns (ODPs) to solve recurrent modeling problems. A specific ODP can be used to derive an Excel template of different terms/annotations and a set of rules that define the relations among those terms/annotations. An Ontorat template is similar to a QTT (Quick Term Template). Such a template can be populated with specific terms or annotations to define or annotate specific ontology terms. With the support of the Ontorat settings, the populated template spreadsheet can then be converted into an OWL file with newly generated ontology terms and axioms.
Reference: Xiang Z, Zheng J, Lin Y, He Y. Ontorat: Automatic generation of new ontology terms, annotations, and axioms based on ontology design patterns. Journal of Biomedical Semantics. 2015, 6:4 doi:10.1186/2041-1480-6-4. PMID: 25785185.
G. Ontobat: Ontology-based data analysis (http://ontobat.hegroup.org)
Ontobat: An Ontology-based biodata analysis tool.
Ontobat is a web-based Semantic Web tool to support ontology-based biological data query and analysis. As a use case, Ontobat includes a program called OntoCOG:, which is an ontology-based statistical tool for analysing the Clusters of Orthologous Groups of proteins (COG).
Unlike other Onto-animal tools, Ontobat focuses on instance level ontology data generation and analysis (Xiang et al., 2014). Ontobat aims to support Linked Open Data (LOD) generation, upload, query, browsing, and statistical analysis. Many features of Ontobat are still under development.
Reference:
H. GOfox: (http://gofox.hegroup.org)
Reference:
I. BFOConvert: (http://bfoconvert.hegroup.org)
Reference:
J. Hegroup RDF triple store (http://sparql.hegroup.org/sparql/)
Most of our ontology tools use our Hegroup Resource Description Framework (RDF) triple store. The SPARQL endpoint of our triple store ...
In summary, the web-based Onto-animal tool package provides a set of comprehensive tools to support ontology development and applications. These tools save time and efforts for ontology developers and users, especially those who do not have or have limited software programming background.
K. Acknowledgements:
The Onto-animal tools were primarily developed in He group at the University of Michigan. The major software developers include Zuoshuang (Allen) Xiang, Bin Zhao, Edison Ong, and Oliver He. Asiyah Yu Lin, a previous postdoc research fellow in He group, contributed a lot by testing the tools with use cases and providing feedbacks. All the source code of these tools were generated in He group. This research was supported by a NIH R01 grant (1R01AI081062).
The development of Onto-animal tools has also obtained supports and collaborations from many collaborators outside the University of Michigan, including Melanie Courtot, Alan Ruttenberg, Chris Mungall, Jie Zheng, among others. The major collaborators of each Onto-animal tools are also co-authors of the Ontoanimal papers.
Your suggestions and comments are welcome. You can directly contact Oliver He (Ontozoo keeper) and send a message through our feedback website. Thank you.
Note: This page is still under construction.
|
